
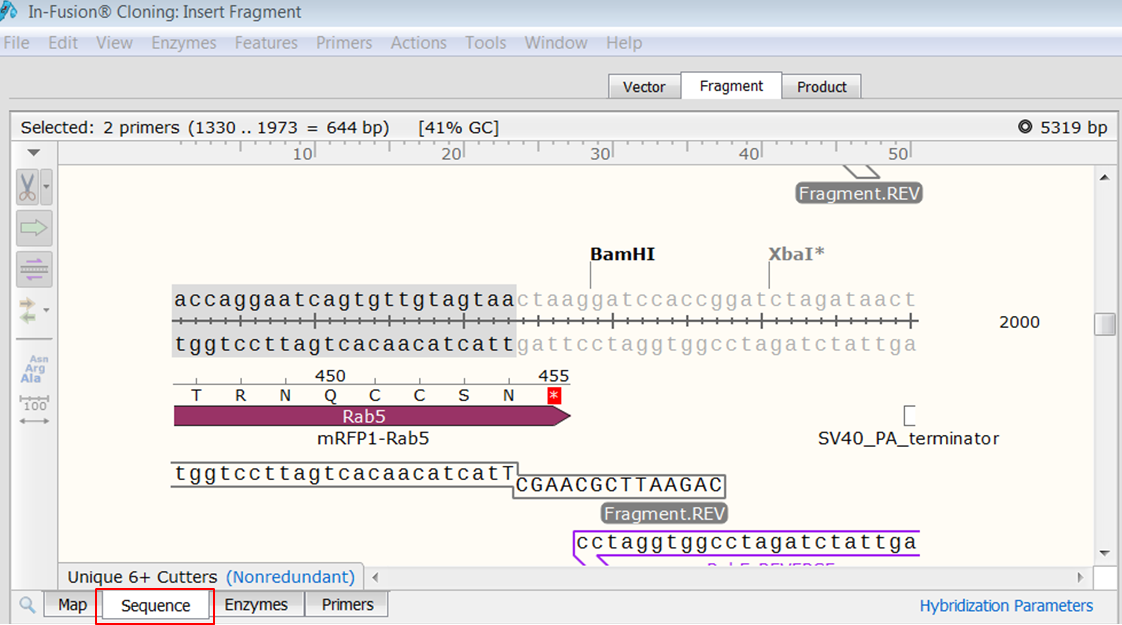
Peptide analysis proceeds by evaluating coverage of the predicted primary protein sequence provided by the initial DNA maps for the VR. To address these problems, we have developed a methodology which combines: 1) selective PCR amplification of VR from both the heavy and light chain IgG from hybridoma, 2) molecular cloning and DNA sequence analysis and 3) tandem mass spectrometry (MS/MS) on enzyme digests obtained from the purified IgG. An incorrect VR sequence will result in a non-functional rAb and de novo assembly of Ig primary structure without a sequence map is challenging. In order to align sequences in SnapGene you should open your sequence and then select Tools-Align Multiple Sequences in the main menu (Figure 3.4.10.1). However the polyploidy nature of a hybridoma cell often results in the added expression of aberrant immunoglobulin-like transcripts or even production of anomalous antibodies which can confound production of rAb. Add homologous sequences to one of the primer pairs to adding all the necessary primers to the same PCR mix. Mutagenesis.with separate primer pairs in the same PCR. forward primer by adding a short sequence complementary to the 5’ end of the reverse primer. This determination can be achieved by sequence analysis of immunoglobulin (Ig) transcripts obtained from a monoclonal antibody (MAb) producing hybridoma and subsequent expression of a rAb. Primer design Using specific primer design (Figure 2), IVA cloning can.
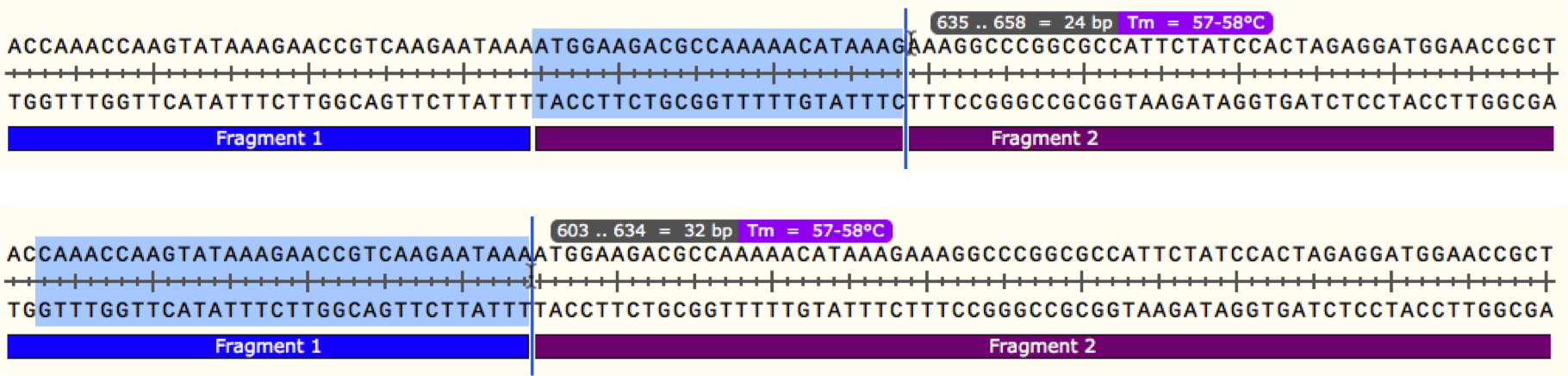
It is the VR that define the unique antigen binding properties and proper sequence identification is essential for functional evaluation and performance of recombinant antibodies (rAb). Four fragments (20 fmol) with 20 bp overlap were assembled using. As your fragment length increases, you should increase the length of the overlapping tail. Overlaps need to anneal efficiently at the reaction temperature of 50C. When induced polymerase chain reaction conditions, the common sequence allows strands from two different fragments to. There are a few general rules for primer design: For simple Gibson assembly reactions, overlapping tails from 15 to 30 nucleotides are sufficient. For a primer binding site, nearest neighbor. An overlap is formed during PCR reaction. For a given duplex, the calculation accounts for any internal mismatches, loops, and dangling ends. This method is the most accurate one available 1,2. Antibody engineering requires the identification of antigen binding domains or variable regions (VR) unique to each antibody. For help designing your primers for use with NEBuilder, please view our primer design. SnapGene calculates oligo melting temperature (Tm) values using a nearest neighbor thermodynamic algorithm with up-to-date parameters 1.
